Julia 笔记¶
- 主页
- My New Workflow with Julia 1.0, mainly focus on how to develop a Julia package.
- Timing in Julia
- Publication quality plots in Julia
not use ~ in serialize
use explicit path.
Version Control¶
- download the “Generic Linux Binaries for x86 (64-bit)” of a particular version from Download Julia
- put it into customed folder, such as
srcunder home directory, - link it to the system path, such as
cd /usr/local/bin
sudo ln -s /home/weiya/src/julia-1.3.1/bin/julia julia1.3.1
Currently, I have installed the following different versions.
$ ll | grep julia
lrwxrwxrwx 1 root root 37 8月 31 2018 julia -> /home/weiya/src/julia-1.0.0/bin/julia*
lrwxrwxrwx 1 root root 37 7月 30 2019 julia1.1.1 -> /home/weiya/src/julia-1.1.1/bin/julia*
lrwxrwxrwx 1 root root 37 9月 18 15:28 julia1.2.0 -> /home/weiya/src/julia-1.2.0/bin/julia*
lrwxrwxrwx 1 root root 37 3月 18 10:25 julia1.3.1 -> /home/weiya/src/julia-1.3.1/bin/julia*
The source folders can be freely moved to another place, and only need to update the symbol links, which can be firstly deleted and then created, or use
sudo ln -sf TARGET LINK_NAME
to override current link. But, if the links is to a folder, then -n need to be added to override,
sudo ln -sfn TARGET LINK_NAME
otherwise, a soft link with the filename of TARGET will be created in the folder LINK_NAME.
since
-f: remove existing destination files-n: treat LINK_NAME as a normal file if it is a symbolic link to a directory
refer to How to change where a symlink points
If LINK_NAME is ., then it would be the same file/folder name as in the TARGET by removing the path, and no need to specify -n for folders.
Also, note that the link to a folder should be deleted with
rm folder
instead of
rm -r folder/ # delete the original folder
Array¶
- drop dimensions:
dropdims, seefor discussion on the necessity to specify the dims to drop
- iterate each row of matrix:
eachrow() - tuple to array:
collect()or[i for i in t] - index of unique elements:
unique(i->x[i], 1:length(x)), used in
pretty println¶
julia> println(a)
[0.4850429731437079 0.1694257866946448; 0.36463804331828364 0.6757027403913352; 0.1855162834299503 0.8954169294007691]
julia> display(a)
3×2 Matrix{Float64}:
0.485043 0.169426
0.364638 0.675703
0.185516 0.895417
More advanced, maybe we can rewrite show to display as you want.
see also: https://stackoverflow.com/questions/35536437/how-to-not-print-types-in-julia
.= vs = when assigning a row/column vector¶
using StatsBase
a = zeros(2, 2)
b = randn(3, 2)
c = mean(b, dims = 1) # 1×2 Array{Float64,2}
a[1, :] = c # OK!
a[:, 1] = c # OK!
a[1, :] .= c # DimensionMismatch("cannot broadcast array to have fewer dimensions")
a[:, 1] .= c # DimensionMismatch("cannot broadcast array to have fewer dimensions")
similarly for column vector,
d = mean(randn(2, 2), dims = 2) # 2×1 Array{Float64,2}
a[1, :] .= d # DimensionMismatch("cannot broadcast array to have fewer dimensions")
one remedy is to reduce the dimension of the row/column vector
a[1, :] = c[:] # OK!
a[1, :] .= c[:] # OK!
an interesting phenomenon is the type returned after assigning regardless a[1,:] or a[:,1]
julia> a[:, 1] = c
1×2 Array{Float64,2}:
julia> a[:, 1] = c[:]
2-element Array{Float64,1}:
julia> a[:, 1] .= c[:]
2-element view(::Array{Float64,2}, :, 1) with eltype Float64:
dims=1¶
总是记不太清 sum, mean 的时候 dims=1 是按行求和还是按列求和,经常先玩个 toy example 才能分辨出来。比如,
julia> a = rand(4, 3)
4×3 Array{Float64,2}:
0.279181 0.0903167 0.148329
0.486691 0.869156 0.0538834
0.110781 0.836284 0.486467
0.810343 0.208659 0.759561
julia> sum(a, dims=1)
1×3 Array{Float64,2}:
1.687 2.00442 1.44824
即把 dims 所代表的维度元素加起来,或者说沿着 dims 进行运算。这一点与 R 的函数 apply 中 margin 参数作用刚好相反,
> a = matrix(rnorm(12), 4, 3)
> a
[,1] [,2] [,3]
[1,] 1.1535199 0.03198768 0.2857126
[2,] -1.3616720 0.32325598 1.9242805
[3,] -0.6519595 -1.14800119 -0.3635221
[4,] 1.2713182 0.98515312 0.2269218
> apply(a, 1, sum)
[1] 1.4712202 0.8858644 -2.1634828 2.4833932
而 Python 的行为与 Julia 类似,先按列求和,再按行求和,但注意其指标从 0 开始,
>>> a = np.random.rand(3, 4);
>>> a
array([[0.47982003, 0.44339329, 0.08865359, 0.30770283],
[0.03171438, 0.82452326, 0.62909043, 0.95733624],
[0.51522304, 0.65602682, 0.95936926, 0.51930784]])
>>> np.sum(a, axis = 0)
array([1.02675744, 1.92394337, 1.67711328, 1.78434691])
>>> np.sum(a, axis = 0, keepdims = True)
array([[1.02675744, 1.92394337, 1.67711328, 1.78434691]])
>>> np.sum(a, axis = 1)
array([1.31956975, 2.44266431, 2.64992695])
>>> np.sum(a, axis = 1, keepdims = True)
array([[1.31956975],
[2.44266431],
[2.64992695]])
另外,Julia 中维度默认不会退化,即对应 Python 中 keepdims = False.
array in functions¶
julia> function g(x)
x += [1, 2]
end
g (generic function with 1 method)
julia> function g!(x)
x .+= [1, 2]
end
g! (generic function with 1 method)
julia> x = [1,2];
julia> g(x)'
1×2 Adjoint{Int64,Array{Int64,1}}:
2 4
julia> x'
1×2 Adjoint{Int64,Array{Int64,1}}:
1 2
julia> g!(x)'
1×2 Adjoint{Int64,Array{Int64,1}}:
2 4
julia> x'
1×2 Adjoint{Int64,Array{Int64,1}}:
2 4
Note that sometimes .+= may make the program much slower, such as en/code/2019-06-14-ML/GD2.jl
no reduction with [1:1]¶
Sometimes, I do not want to the array of array reduces to a single array, then [1:1] would help, instead of [1], see the following toy example.
julia> x
2-element Array{Array{Int64,1},1}:
[1, 2, 3]
[1, 2]
julia> x[1]
3-element Array{Int64,1}:
1
2
3
julia> x[1:1]
1-element Array{Array{Int64,1},1}:
[1, 2, 3]
convert a matrix into an array of array¶
julia> a = zeros(3, 4)
3×4 Array{Float64,2}:
0.0 0.0 0.0 0.0
0.0 0.0 0.0 0.0
0.0 0.0 0.0 0.0
julia> mapslices(x->[x], a, dims = 2)
3×1 Array{Array{Float64,1},2}:
[0.0, 0.0, 0.0, 0.0]
[0.0, 0.0, 0.0, 0.0]
[0.0, 0.0, 0.0, 0.0]
julia> mapslices(x->[x], a, dims = 2)[:]
3-element Array{Array{Float64,1},1}:
[0.0, 0.0, 0.0, 0.0]
[0.0, 0.0, 0.0, 0.0]
[0.0, 0.0, 0.0, 0.0]
refer to
Converting a matrix into an array of arrays
Conversely, we can converting the array of arrays to a matrix, refer to How to convert an array of array into a matrix?
julia> b = mapslices(x->[x], a, dims = 2)[:]
julia> hcat(b...)
4×3 Array{Float64,2}:
0.0 0.0 0.0
0.0 0.0 0.0
0.0 0.0 0.0
0.0 0.0 0.0
julia> reduce(hcat, b)
4×3 Array{Float64,2}:
0.0 0.0 0.0
0.0 0.0 0.0
0.0 0.0 0.0
0.0 0.0 0.0
index from 0¶
using OffsetArrays package, refer to
Base¶
disable warning¶
julia> @warn 1+1
┌ Warning: 2
└ @ Main REPL[1]:1
julia> using Logging
julia> Logging.disable_logging(Logging.Warn)
LogLevel(1001)
julia> @warn 1+1
refers to the official documentation
also tried the command options, -g, --warn-scope, but not work
@__DIR__¶
It expands to the folder of the current file, no matter where we call the script. However, with RCall or PyCall, the expansion would be the folder that executes the script. To expand to the folder of the script, we can first assign it to a variable.
using RCall
using PyCall
println("R: ", rcopy(R"$(@__DIR__)"))
println("Python", py"$(@__DIR__)")
println("Julia:", @__DIR__)
println()
println("Assign @__DIR__ to a variable:")
println()
current_folder = @__DIR__
println("R: ", rcopy(R"$(current_folder)"))
println("Python", py"$(current_folder)")
println("Julia:", current_folder)
Run the above scripts from ~, it outputs
~$ julia1.8 ~/github/techNotes/docs/julia/MWE/dirpath/main.jl
R: /home/weiya
Python/home/weiya
Julia:/home/weiya/github/techNotes/docs/julia/MWE/dirpath
Assign @__DIR__ to a variable:
R: /home/weiya/github/techNotes/docs/julia/MWE/dirpath
Python/home/weiya/github/techNotes/docs/julia/MWE/dirpath
Julia:/home/weiya/github/techNotes/docs/julia/MWE/dirpath
Relative path in include vs read¶
Suppose we have two files
# file: relpath/a.jl
f(x) = x + 1
and
# file: relpath/a.jl
include("a.jl")
println(f(1))
println(String(read("a.jl")))
If we run the scripts from docs/julia/MWE
# file: relpath/b.jl
$ julia relpath/b.jl
it will throw an error that a.jl is not found. But note that include also use the same relative path. The reason is that
help?> include
...During including, a task-local include path is set to the
directory containing the file.
连等号赋值¶
如果采用 a=b=c=ones(10) 形式赋值的话,则如果后面改变 a 的值,b 和 c 的值也将随之改变。
但如果 a=b=c=1 为常值的话,则三个变量的值还是独立的。
ERROR: expected Type{T}¶
参考 ERROR: LoadError: TypeError: Type{…} expression: expected Type{T}, got Module
其中举了一个小例子
module Foo end
Foo{Int64}
会爆出这样的错误。但是一开始竟然没有仔细类比,最后在 REPL 中逐行试验才发现是,using SharedArrays 后直接用 SharedArrays{Float64}(10),这与上面 Foo 的错误形式完全一样,竟然没有仔细类比。哎,看来以后多思考一下错误可能的原因,不要一味蛮力试验。
type, instance, and object¶
看两个句子:
- A type union is a special abstract type which includes as objects all instances of any of its argument types
Nothingis the singleton type whose only instance is the objectnothing.
从中分析知道,instance 相对于 types,而 object 相对 instance。一个 type 可能有多个 instance,每个 instance 称之为 object。
- instance of some types
- object of some instances
Control Flow¶
priority of &¶
use
(x[1] <= width) & (x[1] >= 0) & (x[2] <= height) & (x[2] >=0)
instead of
x[1] <= width & x[1] >= 0 & x[2] <= height & x[2] >=0
ifelse vs ?¶
ifelse differs from ? or if in that it is an ordinary function, so all the arguments are evaluated first.
For example, it will try to evaluate the second clause,
julia> isnothing(nothing) ? "" : @sprintf "%.4f" nothing
""
julia> ifelse(isnothing(nothing), "" , @sprintf "%.4f" nothing)
ERROR: MethodError: no method matching isfinite(::Nothing)
Adopted from Clouds#32
Tip
The specifiers in @sprintf should be consists of ones in C++, and the detailed explanation refers to Print Format – C/C++
Note that, %g uses the shorter representation between %e and %f.
for in array¶
julia> for i = 1:3
a = [i for i = i:3]
println(a)
end
[1, 2, 3]
[2, 3]
[3]
same iterative variable i, but it works well.
Distributed¶
@distributed¶
如果配合 sharedarrays 使用时,需要加上 @sync, 参考@fetch
@sync vs @async¶
julia> @time sleep(2)
2.003579 seconds (5 allocations: 128 bytes)
julia> @time @async sleep(2)
0.015021 seconds (6.57 k allocations: 379.004 KiB)
Task (runnable) @0x00007f24428409d0
julia> @time @sync @async sleep(2)
2.008425 seconds (1.62 k allocations: 79.125 KiB)
Task (done) @0x00007f24429ebd00
@async: for whatever falls within its scope, Julia will start this task running but then proceed to whatever comes next in the script without waiting for the task to be complete@sync: wait until all lexically-enclosed uses of@async,@spawn,@spawnatand@distributedare complete, where complete matters how you define the tasks within the scope, such asremotecall_fetch(only finished when it gets the message from the worker that its task is complete) vsremotecall(finished once it has sent the worker the job to do)
With setting
using Distributed
cell(N) = Vector{Any}(undef, N)
addprocs(2)
julia> @time begin
a = cell(nworkers())
for (idx, pid) in enumerate(workers())
a[idx] = remotecall_fetch(sleep, pid, 2)
end
end
4.270026 seconds (28.35 k allocations: 1.384 MiB)
julia> @time begin
a = cell(nworkers())
@sync for (idx, pid) in enumerate(workers())
@async a[idx] = remotecall_fetch(sleep, pid, 2)
end
end
2.031801 seconds (2.50 k allocations: 144.907 KiB)
julia> @time begin
a = cell(nworkers())
for (idx, pid) in enumerate(workers())
println("sending work to $pid")
@async a[idx] = remotecall_fetch(sleep, pid, 2)
end
end
sending work to 2
sending work to 3
0.020147 seconds (2.10 k allocations: 121.017 KiB)
julia> a
2-element Array{Any,1}:
#undef
#undef
# wait around 2 seconds
julia> a
2-element Array{Any,1}:
nothing
nothing
julia> @time begin
a = cell(nworkers())
@async for (idx, pid) in enumerate(workers())
println("sending work to $pid")
a[idx] = remotecall_fetch(sleep, pid, 2)
end
end
sending work to 2
0.000141 seconds (32 allocations: 2.867 KiB)
Task (runnable) @0x00007f2442843d00
# wait around 2 seconds, it prints
julia> sending work to 3
julia> @time begin
a = cell(nworkers())
@sync @async for (idx, pid) in enumerate(workers())
a[idx] = remotecall_fetch(sleep, pid, 2)
end
end
4.017451 seconds (13.96 k allocations: 692.136 KiB)
Task (done) @0x00007f2442dfdfc0
refer to How and When to Use @async and @sync in Julia - Stack Overflow
And as the official documentation said,
@async is similar to @spawnat, but only runs tasks on the local process. We use it to create a “feeder” task for each process. Each task picks the next index that needs to be computed, then waits for its process to finish, then repeats until we run out of indices. Note that the feeder tasks do not begin to execute until the main task reaches the end of the @sync block, at which point it surrenders control and waits for all the local tasks to complete before returning from the function.
Info
Applications in my project: parallel.job, where it is much like the feeder, although it might be quite fast.
Gurobi¶
- Currently (2022-09-23) no way to enable Gurobi.jl in GitHub Actions.
- hide message of
Set parameter ...:GRBsetintparam(GRB_ENV, GRB_INT_PAR_OUTPUTFLAG, 0). Inspired by the way for C++ API of Gurobi,and
JLD vs JLD2¶
JLD depends on HDF5, whose installation requires to login in, while JLD2 avoid the dependence of HDF5.
Juno¶
Info
An IDE on Atom.
Shortcuts¶
Ctrl+J Ctrl+E: switch from REPL to editor, refer to the correct shortcut is be Ctrl-J Ctrl-E (and the command is called Julia Client: Focus Last Editor), an inspiration is that I can check it viaCtrl-Shift-P.
Bug when Cltr + Enter¶
当我使用 Ctrl + Enter 运行下列语句时,
using Images, ImageView
img = load(download("https://juliaimages.org/latest/assets/segmentation/horse.jpg"));
imshow(img)
REPL 冻住了,一直停在
Dict{String,Any} with 4 entries:
而我如果直接将上述语句复制到 REPL 中运行时,一切 OK,正常显示为
Dict{String,Any} with 4 entries:
"gui" => Dict{String,Any}("window"=>GtkWindowLeaf(name="", parent, wi…
"roi" => Dict{String,Any}("redraw"=>50: "map(clim-mapped image, input…
"annotations" => 3: "input-2" = Dict{UInt64,Any}() Dict{UInt64,Any}
"clim" => 2: "CLim" = CLim{RGB{Float64}}(RGB{Float64}(0.0,0.0,0.0), RG…
找到类似的 Issue: [BUG] Evaluation in Editor freezes Atom occasionally
虽然其解决方案不太清楚,是有关 notification-daemon,但是维护者 @pfitzseb 的回复
OS level notifications are disabled completly in the latest Juno release because they were causing crashes on Mac and didn’t really seem to work on other platforms.
提醒我或许是因为 Juno 的版本问题,当前版本为 v0.7.2, 最新版本为 v0.8.1
仅仅更新 Juno.jl 的版本似乎还不行,另外将 Atom.jl 从 v0.12.8 更新到 v0.12.9
最后竟然真的解决了!
System Proxy¶
当在 .bashrc 添加 http_proxy 和 https_proxy 中并 source 之后,直接在 shell 里面开的 julia session 中,验证当前 ip
run(`curl ifconfig.me`)
或者直接查看
ENV["http_proxy"]
发现可以使用系统代理。但是在 Atom 中开的 julia,则不行。找到 Atom 一条相关的 issue: [Atom 1.32.0] Does not inherit environment variables when launched not from the command line,但是我已经是最新版了,不应该存在这个问题。但是当我通过 Ctrl-Shift-i 打开 devtools,验证 process.env["http_proxy"],显示 undefined;而且直接在 source 之后的 terminal 中打开,也显示 undefined。猜想,只对 PATH 有效?或者因为 Juno.jl?
跳过这个问题,最后直接在 startup.jl 文件中添加
ENV["http_proxy"] =
ENV["https_proxy"]
进行设置,这参考了 Install packages behind the proxy
Function¶
calculation in arguments¶
julia> f(x; n = 10, m=n/2) = x+m
f (generic function with 1 method)
julia> f(1)
6.0
where m = n/2 is allowed. But Python does not support this feature.
Bug
>>> def f(x, n = 10, m = n/2):
... x + m
...
Traceback (most recent call last):
File "<stdin>", line 1, in <module>
NameError: name 'n' is not defined
function redefinition in if¶
julia> function f()
flag = true
if flag
g(x) = 1
else
g(x) = 2
end
println(g(1))
end
WARNING: Method definition g(Any) in module Main at REPL[1]:4 overwritten at REPL[1]:6.
f (generic function with 1 method)
julia> f()
2
julia> function f()
flag = true
if flag
g = x->1
else
g = x->2
end
println(g(1))
end
f (generic function with 1 method)
julia> f()
1
The correct way is to use the anonymous function, see also Defining a function inside if…else..end NOT as expected? - New to Julia - JuliaLang
invoke function via string¶
julia> getfield(Main, Symbol("exp"))(1)
2.718281828459045
refer to reflection - Julia: invoke a function by a given string - Stack Overflow
default argument and optional arguments¶
julia> function f(x, y=nothing; z=1)
if isnothing(y)
println("x + z = $(x+z)")
else
println("x + y + z = $(x+y+z)")
end
end
julia> f(1)
x + z = 2
julia> f(1, 2)
x + y + z = 4
with this format, we can avoid define two functions f(x; ) and f(x, y;) respectively.
function in function¶
julia> function f2()
a = 10
function g()
a = a+10
end
g()
println(a)
end
f2 (generic function with 1 method)
julia> f2()
20
but
julia> function f2()
a = 10
g() = a + 10
g()
println(a)
end
f2 (generic function with 1 method)
julia> f2()
10
since the first one does not have return, but g() has return and the addition is not on a.
Pkg¶
- Pkg + BinaryBuilder: binary objects that are not Julia packages
- [
dev+test]: test project instead ofincludescripts. Specifically, open a Julia session without--projectoption in the project folder, and then switch into the Pkg environment, typedev .&test YourProjectName. See more attempts.
PyCall¶
Update Matplotlib¶
Info
Post: 2022-04-09 19:13:48
julia> pyplot()
[ Info: Precompiling PyPlot [d330b81b-6aea-500a-939a-2ce795aea3ee]
┌ Warning: You are using Matplotlib 3.1.3, which is no longer
│ officialy supported by the Plots community. To ensure smooth Plots.jl
│ integration update your Matplotlib library to a version >= 3.4.0
│
│ If you have used Conda.jl to install PyPlot (default installation),
│ upgrade your matplotlib via Conda.jl and rebuild the PyPlot.
│
│ If you are not sure, here are the default instructions:
│
│ In Julia REPL:
│ ```
│ import Pkg;
│ Pkg.add("Conda")
│ import Conda
│ Conda.update()
│ Pkg.build("PyPlot")
│ ```
│
└ @ Plots ~/.julia/packages/Plots/9C6z9/src/backends/pyplot.jl:29
Then follow the instruction,
julia> import Conda
julia> Conda.update()
[ Info: Running `conda update -y --all conda` in root environment
Collecting package metadata (current_repodata.json): done
Solving environment: done
## Package Plan ##
environment location: /home/weiya/.julia/conda/3
added / updated specs:
- conda
it implies that the env created by Conda.jl is ~/.julia/conda/3
If I check all env in the shell, it returns
$ conda env list
# conda environments:
#
/home/weiya/.julia/conda/3
/home/weiya/.julia/conda/3/envs/_ORCA_jl_
base /home/weiya/anaconda3
py37 /home/weiya/anaconda3/envs/py37
...
Couldn’t find libpython error¶
ENV["PYTHON"]=""; Pkg.build("PyCall")
to install its own private Miniconda distribution for you, in a way that won’t interfere with your other Python installations.
Refer to Couldn’t find libpython error #199
Actually, using PyPlot also encounters the similar question, cannot find matplotlib, then I changed to the conda environment contained such package, and then
ENV["PYTHON"]="the-python-path";
# change to package command line
build PyCall
it works. And it seems that it does NOT depend on conda environment any more, i.e., PyPlot still works for different conda environment, even without matplotlib package in such environment.
In the words of the official documentation,
If you use Python virtualenvs, then be aware that PyCall uses the virtualenv it was built with by default, even if you switch virtualenvs. If you want to switch PyCall to use a different virtualenv, then you should switch virtualenvs and run rm(Pkg.dir(“PyCall”,”deps”,”PYTHON”)); Pkg.build(“PyCall”).
which supports my above guess.
Record for install PyCall¶
Switch to py37 conda environment, then run julia1.2.0
(v1.2) pkg> add PyCall
Then test to import module in the same REPL,
julia> using PyCall
[ Info: Recompiling stale cache file /home/weiya/.julia/compiled/v1.2/PyCall/GkzkC.ji for PyCall [438e738f-606a-5dbb-bf0a-cddfbfd45ab0]
julia> math = pyimport("math")
PyObject <module 'math' from '/home/weiya/anaconda3/lib/python3.7/lib-dynload/math.cpython-37m-x86_64-linux-gnu.so'>
Now open a new julia session under different conda environment (or no any conda environment), re-run the above code
julia> using PyCall
julia> math = pyimport("math")
PyObject <module 'math' from '/home/weiya/anaconda3/lib/python3.7/lib-dynload/math.cpython-37m-x86_64-linux-gnu.so'>
As you can see, no any compiling info when using PyCall, and return the same math module. However such module does not corresponds to the py37 conda environment. The official documentation says
On GNU/Linux systems, PyCall will default to using the python3 program (if any, otherwise
python) in your PATH.
Then to specify the version of python such as
julia> ENV["PYTHON"] = Sys.which("python")
"/home/weiya/anaconda3/envs/py37/bin/python3.7"
and rebuild, note that need to restart the julia session to recompiling,
julia> using PyCall
[ Info: Recompiling stale cache file /home/weiya/.julia/compiled/v1.2/PyCall/GkzkC.ji for PyCall [438e738f-606a-5dbb-bf0a-cddfbfd45ab0]
julia> pyimport("math")
PyObject <module 'math' from '/home/weiya/anaconda3/envs/py37/lib/python3.7/lib-dynload/math.cpython-37m-x86_64-linux-gnu.so'>
then the module corresponds to the specified conda environment, and then like the above test, it also can be used in other conda environment without recompiling.
RCall¶
julia> rcopy(R"lm($dop ~ 0 + $H)$coefficients")
100-element reshape(::Array{Union{Missing, Float64},1}, 100) with eltype Union{Missing, Float64}:
missing
-1.909321682273283e-23
A related error is
julia> Array{Float64}(rcopy(R"lm($dop ~ 0 + $H)$coefficients"))
ERROR: MethodError: Cannot `convert` an object of type Missing to an object of type Float64
Closest candidates are:
convert(::Type{T}, ::T) where T<:Number at number.jl:6
convert(::Type{T}, ::Number) where T<:Number at number.jl:7
convert(::Type{T}, ::Base.TwicePrecision) where T<:Number at twiceprecision.jl:250
Random¶
Case 1: Two files¶
Suppose I have the following two files,
using Random
import StatsBase.sample
function g(seed = 1234)
println(sample(Random.seed!(seed), 1:5, 5))
end
include("a.jl")
function f()
println(rand(3))
g(1234)
println(rand(3))
end
after first different results, it repeats to show the same results.
julia> include("b.jl")
f (generic function with 1 method)
julia> f()
[0.8484503038106239, 0.6270935436096212, 0.8958165220379257]
[3, 4, 5, 4, 1]
[0.3728458178193992, 0.2631210819875911, 0.9886904946486306]
julia> f()
[0.4898584067326506, 0.4252112774882528, 0.37976500269509095]
[3, 4, 5, 4, 1]
[0.3728458178193992, 0.2631210819875911, 0.9886904946486306]
julia> f()
[0.4898584067326506, 0.4252112774882528, 0.37976500269509095]
[3, 4, 5, 4, 1]
[0.3728458178193992, 0.2631210819875911, 0.9886904946486306]
Case 2: a single file¶
The above phenomenon can be observed in a single file,
using Random
function f(seed)
println(rand(Random.seed!(seed)))
end
function g(seed)
println(rand())
f(seed)
println(rand())
end
julia> include("c.jl")
g (generic function with 1 method)
julia> g(1)
0.8797616728308622
0.23603334566204692
0.34651701419196046
julia> g(1)
0.3127069683360675
0.23603334566204692
0.34651701419196046
julia> g(1)
0.3127069683360675
0.23603334566204692
0.34651701419196046
A Correct Way¶
replace Random.seed!(seed) with MersenneTwister
using Random
function f(seed)
println(rand(MersenneTwister(seed)))
end
function g(seed)
println(rand())
f(seed)
println(rand())
end
it behaves as expected,
julia> include("d.jl")
g (generic function with 1 method)
julia> g(1)
0.12495220703448995
0.23603334566204692
0.41171274946529546
julia> g(1)
0.7243446710747494
0.23603334566204692
0.3672395293576307
julia> g(1)
0.5368102592873767
0.23603334566204692
0.6466136938696847
Revise¶
Is there a way to undo using in Julia?¶
NO
- https://stackoverflow.com/questions/33927523/can-i-make-julia-forget-a-method-from-the-repl
- https://stackoverflow.com/questions/36249313/is-there-a-way-to-undo-using-in-julia
Although we cannot remove some defined function, with the powerful Revise.jl, we can update the functions without reloading the scripts.
Statistics (StatsBase, Distributions)¶
关于 mean()¶
using Statistics后才能用mean(),而using Distributions后也能用mean()。前者表示generic function with 5 methods,后者称generic function with 78 methods.
Normal 中标准差为 0 的问题¶
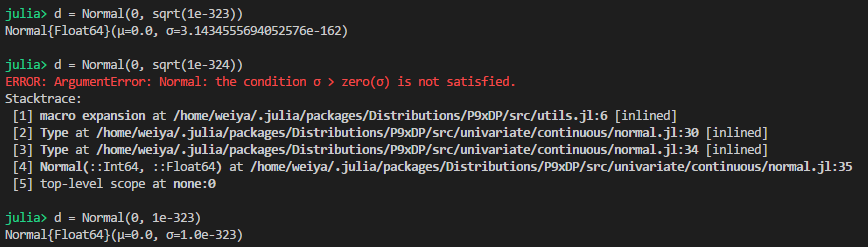
可知,最低可以支持 1e-323,所以似乎也支持 sqrt(1e-646),但并没有,而且当 sqrt(1e-324) 时精度就不够了,似乎 sqrt(x) 的精度与 x 的精度相当。
SymPy: Symbolic Math¶
julia> using SymPy
[ Info: Precompiling SymPy [24249f21-da20-56a4-8eb1-6a02cf4ae2e6]
ERROR: InitError: PyError (PyImport_ImportModule
The Python package sympy could not be imported by pyimport. Usually this means
that you did not install sympy in the Python version being used by PyCall.
PyCall is currently configured to use the Python version at:
/home/weiya/anaconda3/envs/py37/bin/python3.7
and you should use whatever mechanism you usually use (apt-get, pip, conda,
etcetera) to install the Python package containing the sympy module.
julia> @vars x
(x,)
julia> exp(-x^2/2)/sqrt(2pi)
2
-x
────
2
0.398942280401433⋅ℯ
julia> f(x) = exp(-x^2/2)/sqrt(2pi)
f (generic function with 1 method)
julia> integrate(f(x), x)
⎛√2⋅x⎞
0.199471140200716⋅√2⋅√π⋅erf⎜────⎟
⎝ 2 ⎠
julia> integrate(f(x), (x, -Inf, Inf))
0.398942280401433⋅√2⋅√π
julia> integrate(x*f(x), (x, -Inf, Inf))
0
julia> integrate(x*f(x), x)
2
-x
────
2
-0.398942280401433⋅ℯ
julia> integrate(x*f(x), (x, -Inf, Inf))
0
julia> integrate(f(x)^2, (x, -Inf, Inf))
0.159154943091895⋅√π
julia> integrate(f(x)^2, x)
0.0795774715459477⋅√π⋅erf(x)
where erf is https://en.wikipedia.org/wiki/Error_function
parallel¶
references
- Julia parallel computing over multiple nodes in cluster
- Using julia -L startupfile.jl, rather than machinefiles for starting workers.
- Help setting up Julia on a cluster
pbsdsh (unsolved)¶
Submit a pbsdsh job and specify multiply nodes with multiply cores, say nodes=2:ppn=4
error log file (see full file in the cluster: dshmcmc.e21907):
fatal: error thrown and no exception handler available.
InitError(mod=:Base, error=ArgumentError(msg="Package Sockets not found in current path:
- Run `Pkg.add("Sockets")` to install the Sockets package.
"))
rec_backtrace at /buildworker/worker/package_linux64/build/src/stackwalk.c:94
record_backtrace at /buildworker/worker/package_linux64/build/src/task.c:246
jl_throw at /buildworker/worker/package_linux64/build/src/task.c:577
require at ./loading.jl:817
init_stdio at ./stream.jl:237
jfptr_init_stdio_4446.clone_1 at /opt/share/julia-1.0.0/lib/julia/sys.so (unknown line)
jl_apply_generic at /buildworker/worker/package_linux64/build/src/gf.c:2182
reinit_stdio at ./libuv.jl:121
__init__ at ./sysimg.jl:470
jl_apply_generic at /buildworker/worker/package_linux64/build/src/gf.c:2182
jl_apply at /buildworker/worker/package_linux64/build/src/julia.h:1536 [inlined]
jl_module_run_initializer at /buildworker/worker/package_linux64/build/src/toplevel.c:90
_julia_init at /buildworker/worker/package_linux64/build/src/init.c:811
julia_init__threading at /buildworker/worker/package_linux64/build/src/task.c:302
main at /buildworker/worker/package_linux64/build/ui/repl.c:227
__libc_start_main at /lib64/libc.so.6 (unknown line)
_start at /opt/share/julia-1.0.0/bin/julia (unknown line)
But when I just use single node, and arbitrary cores, say nodes=1:ppn=2, it works well.
references¶
julia local package 失败折腾记录¶
- no error after
add ~/GitHub/adm.jl, butusing admcannot work. refer to Adding a local package - set
startup.jlbut still not work. refer to How does Julia find a module? - one possible way: Finalizing Your Julia Package: Documentation, Testing, Coverage, and Publishing
Variable Scopes¶
for loop¶
i = 0
for j = 1:2
if j != i
println("ok")
end
end
i = 0
for j = 1:2
if j != i
println("ok")
end
i = j
end
i = 0
for j = 1:2
if j != i
println("ok")
end
global i = j
end
The results are
$ julia mwe1.jl
ok
ok
$ julia mwe2.jl
ERROR: LoadError: UndefVarError: i not defined
# more information given in julia1.5.2
$ julia1.5.2 mwe2.jl
┌ Warning: Assignment to `i` in soft scope is ambiguous because a global variable by the same name exists: `i` will be treated as a new local. Disambiguate by using `local i` to suppress this warning or `global i` to assign to the existing global variable.
ERROR: LoadError: UndefVarError: i not defined
$ julia mwe3.jl
ok
ok
Thus, the reason is that i is a global variable, we cannot modify a global variable in a local block without global keyword, i.e., global i = i + 1, but we can read i in the for loop.
Alternatively, we can use let block,
let
i = 0
for j = 1:2
i = i + 1
end
i
end
then i isn’t really a global variable anymore.
References:
while loop¶
function test_while(flag)
if flag
a = 1
else
while true
a = 2
break
end
end
println(a)
end
function test_while2(flag)
while true
a = 2
break
end
println(a)
end
the results are
julia> test_while(false)
2
julia> test_while2(false)
ERROR: UndefVarError: a not defined
自定义 == and hash()¶
对于自定义的 mutable struct, 直接用 == 返回 false,因此需要自己定义等号,比如
Base.:(==)(x::Reaction, y::Reaction) = all([
x.substrate == y.substrate,
x.product == y.product,
x.reversible == y.reversible,
x.species == y.species])
而如果想用 unique 函数的话,其基于 isequal 函数,而 isequal 是通过判断 hash 值来判定的,所以仅仅定义了 == 仍不够,还需要定义 hash 函数,这很简单,比如
function Base.hash(obj::Reaction, h::UInt)
return hash((obj.substrate, obj.product, obj.reversible, obj.species), h)
end
hash(散列、杂凑)函数,是将任意长度的数据映射到有限长度的域上。直观解释起来,就是对一串数据m进行杂糅,输出另一段固定长度的数据h,作为这段数据的特征(指纹)。
or
HASH函数是这么一种函数,他接受一段数据作为输入,然后生成一串数据作为输出,从理论上说,设计良好的HASH函数,对于任何不同的输入数据,都应该以极高的概率生成不同的输出数据,因此可以作为“指纹”使用,来判断两个文件是否相同。 数据 ---->输入 HASH 函数 ---->输出指纹数据
参考
- Hash function for custom type
- Surprising struct equality test
- What is the difference between “using” and “import”?
import vs using
Instead of explicitly redefine function with Module.function name, we can first import it, e.g., pdf, we need to use import Distributions.pdf.
MethodError: objects of type Module are not callable¶
check if the function is mistaken by the module name, such as AxisArray vs. AxisArrays.
/lib/x86_64-linux-gnu/libz.so.1: versionZLIB_1.2.9’ not found`¶
work with GPU¶
- An Introduction to GPU Programming in Julia
- Re-build after install cudnn, refer to GPU on cluster: conversion to pointer not defined for CuArray{Float32,2}
load ImageView on Server¶
when using ImageView, it throws an error,
ERROR: LoadError: InitError: Cannot open display:
fixed by ssh -X. If further ssh on the node, it still works.
-i vs -L¶
if need to REPL and also arguments, then -i would be suitable.
And note that -- should be separated the switches and program files. (but seems not necessary)
Unable to display plot using the REPL. GKS errors¶
The reason would be relate to the GR package,
cd .julia/packages/GR/ZI5OE/deps/gr/bin/
ldd gksqt
and then get
linux-vdso.so.1 => (0x00007ffdc1e58000)
libQt5Widgets.so.5 => not found
libQt5Gui.so.5 => not found
libQt5Network.so.5 => not found
libQt5Core.so.5 => not found
libGL.so.1 => /lib64/libGL.so.1 (0x00002ae608a73000)
libpthread.so.0 => /lib64/libpthread.so.0 (0x00002ae608ce5000)
libstdc++.so.6 => /lib64/libstdc++.so.6 (0x00002ae608f01000)
libm.so.6 => /lib64/libm.so.6 (0x00002ae609209000)
libgcc_s.so.1 => /lib64/libgcc_s.so.1 (0x00002ae60950b000)
libc.so.6 => /lib64/libc.so.6 (0x00002ae609721000)
libexpat.so.1 => /lib64/libexpat.so.1 (0x00002ae609aef000)
libxcb-dri3.so.0 => /lib64/libxcb-dri3.so.0 (0x00002ae609d19000)
libxcb-xfixes.so.0 => /lib64/libxcb-xfixes.so.0 (0x00002ae609f1c000)
libxcb-present.so.0 => /lib64/libxcb-present.so.0 (0x00002ae60a125000)
libxcb-sync.so.1 => /lib64/libxcb-sync.so.1 (0x00002ae60a328000)
libxshmfence.so.1 => /lib64/libxshmfence.so.1 (0x00002ae60a52f000)
libglapi.so.0 => /lib64/libglapi.so.0 (0x00002ae60a733000)
libselinux.so.1 => /lib64/libselinux.so.1 (0x00002ae60a963000)
libXext.so.6 => /lib64/libXext.so.6 (0x00002ae60ab8a000)
libXdamage.so.1 => /lib64/libXdamage.so.1 (0x00002ae60ad9d000)
libXfixes.so.3 => /lib64/libXfixes.so.3 (0x00002ae60afa0000)
libX11-xcb.so.1 => /lib64/libX11-xcb.so.1 (0x00002ae60b1a6000)
libX11.so.6 => /lib64/libX11.so.6 (0x00002ae60b3a9000)
libxcb.so.1 => /lib64/libxcb.so.1 (0x00002ae60b6e7000)
libxcb-glx.so.0 => /lib64/libxcb-glx.so.0 (0x00002ae60b90f000)
libxcb-dri2.so.0 => /lib64/libxcb-dri2.so.0 (0x00002ae60bb2b000)
libXxf86vm.so.1 => /lib64/libXxf86vm.so.1 (0x00002ae60bd30000)
libdrm.so.2 => /lib64/libdrm.so.2 (0x00002ae60bf36000)
libdl.so.2 => /lib64/libdl.so.2 (0x00002ae60c148000)
/lib64/ld-linux-x86-64.so.2 (0x0000557abc358000)
libXau.so.6 => /lib64/libXau.so.6 (0x00002ae60c34c000)
libpcre.so.1 => /lib64/libpcre.so.1 (0x00002ae60c551000)
(refer to one comment in Error: GKS: can’t connect to GKS socket application)
So it seems that it is due to the missing of libqt5. If I have the sudo privilege, maybe just type
sudo apt-get install qt5-default
but I cannot. But I noticed that I have installed miniconda3, and the qt is installed,
qmake -v
# QMake version 3.1
# Using Qt version 5.9.7 in /users/xxx/miniconda3/lib
So a natural way is to append the library path of qt to LD_LIBRARY_PATH, that is,
# in .bashrc
export LD_LIBRARY_PATH=/users/xxx/miniconda3/lib:$LD_LIBRARY_PATH
then rebuild GR package in the julia REPL.
This solution should work, but actually it failed at the first time. Do not be too frustrated, I found the reason is that I type miniconda3 as minconda3 in the path.
It works now, although it still throw an error,
julia> libGL error: unable to load driver: swrast_dri.so
libGL error: failed to load driver: swrast
reset kw... value¶
In the following situation, I want to reset a particular argument in kw....
function f(; kw...)
g(; kw...)
end
Note that the type of kw is Base.Iterators.Pairs, firstly I want to directly reset the value via
h(;kw...) = kw
h(a=1).data.a = 2
but it throws
ERROR: setfield! immutable struct of type NamedTuple cannot be changed
Stacktrace:
[1] setproperty!(::NamedTuple{(:a,),Tuple{Int64}}, ::Symbol, ::Int64) at ./Base.jl:21
[2] top-level scope at none:0
So I need to find new method, or give up such idea. Fortunately, the official documentation of Julia says
The nature of keyword arguments makes it possible to specify the same argument more than once. For example, in the call
plot(x, y; options..., width=2)it is possible that the options structure also contains a value forwidth.
Thus, I can write
function f(; kw...)
g(; kw..., b = 6)
end
b is the argument I want to reset.
some interesting behavior¶
d = Dict{Int, Int}()
d[1] = 1
if we access
d.vals
# d.keys
it will return a 16-element Array{Int64,1}, but if we use
values(d)
keys(d)
it correctly returns the exactly one elements, but with type Base.ValueIterator for a Dict{Int64,Int64} with 1 entry.
pass array into function¶
julia> a = [1,2]
2-element Array{Int64,1}:
1
2
julia> function f(a)
a = a .+ 1
end
f (generic function with 1 method)
julia> f(a)
2-element Array{Int64,1}:
2
3
julia> a
2-element Array{Int64,1}:
1
2
julia> function g(a)
a .= a .+ 1
end
g (generic function with 1 method)
julia> g(a)
2-element Array{Int64,1}:
2
3
julia> a
2-element Array{Int64,1}:
2
3
check whether array entry is undef¶
b = Array{Array{Int, 1}}(undef, 3)
isdefined(b, 1)
# or
isassigned(b, 1)
# not
# isdefined(b[1])
refer to Julia: check whether array entry is undef
Memory Allocation¶
Usage of GC.gc()¶
Manually run GC.gc() can release the occupied memory,
The experiments are as follows,
- open a
julia1.6session - run
tseries = zeros(1024,1024, 5, 50);, it allocates 1.953GB, roughtly1.953/16 = 12%, while the columnMEM%displayed byhtopindeed show11.4. - repeat to run, then
MEM%increses to21.5 - run
GC.gc(), thenMEM%decreases to11.0 - repeat to run 2 times, then
MEM%increases to31.2 - run
GC.gc(), thenMEM%decreases to11.0 - set
tseries = nothing, and runGC.gc(), it disappears from the top line ofhtop, i.e., (nearly) zero memory.
Refer to the following video for the whole experiments.
Another experiments are for for loop,
julia> for i = 1:6
tseries = zeros(1024,1024, 5, 50);
sleep(3)
if i % 3 == 0
println("gc...")
GC.gc(); sleep(3)
end
end
It can be observed that MEM% increses from ~10 to ~20, and then ~30, then decreases to ~10 after first GC.
With monitoring the memory from top, we can use the following memuse() function to get the memory usage of the current process.
function memuse()
pid = getpid()
return round(Int,parse(Int,read(`ps -p $pid -o rss=`, String))/1024)
end
println("init mem = ", memuse(), "MB")
for i = 1:6
# tseries = zeros(1024,1024, 5, 50);
r = R"rnorm(1024*1024*5*50)"
# sleep(3)
println("i = ", i, ", mem = ", memuse(), "MB")
if i % 3 == 0
finalize(r)
GC.gc(); GC.gc(); GC.gc(); GC.gc();
R"gc()"; R"gc()";
sleep(3)
println("after gc..., mem = ", memuse(), "MB")
end
end
function test()
for i = 1:2 println("\ni=$i")
a = rand(10000,10000)
println("Created a $(memuse())")
a = 0
GC.gc()
println("Release a $(memuse())\n")
b = rand(10000,10000)
println("Created b $(memuse())")
b = 0
GC.gc()
println("Release b $(memuse())\n")
c = rand(10000,10000)
println("Created c $(memuse())")
c =0
GC.gc()
println("Release c $(memuse())\n")
end
end
See also:
Note
The most possible reason for abnormally increasing memory is due to memory leak instead of insufficient gc(). This part is my exploration when I encounter (possibly) memory leak with ECOS, but the issue disappears when I switched it to Gurobi. Refer to this private commit for more details.
memory allocation of undef¶
julia> @time Array{Array{Int, 1}, 2}(undef, 100, 100);
0.000024 seconds (6 allocations: 78.359 KiB)
julia> @time zeros(100,100, 1);
0.000014 seconds (6 allocations: 78.359 KiB)
julia> @time zeros(100,100, 10);
0.000077 seconds (6 allocations: 781.484 KiB)
wrong arrangement of multiple figures in a grid¶
Hi, I am confused by the arrangement of multiple figures in a grid. I begin with the example code in the README,
gui = imshow_gui((300, 300), (2, 1)) # 2 columns, 1 row of images (each initially 300×300)
canvases = gui["canvas"]
imshow(canvases[1,1], testimage("lighthouse"))
imshow(canvases[1,2], testimage("mandrill"))
Gtk.showall(gui["window"])
it throws an error when accessing canvases[1,2]
julia> imshow(canvases[1,2], testimage("mandrill"))
ERROR: BoundsError: attempt to access 2×1 Array{Any,2} at index [1, 2]
and I check the size of canvases is
julia> size(canvases)
(2, 1)
it seems that the canvases will arrage by column while the gui declares it should be by rows.
On the other hand, if I run
gui = imshow_gui((300, 300), (2, 1)) # 2 columns, 1 row of images (each initially 300×300)
canvases = gui["canvas"]
imshow(canvases[1,1], testimage("lighthouse"))
imshow(canvases[2,1], testimage("mandrill"))
Gtk.showall(gui["window"])
gui = imshow_gui((300, 300), (1, 2)) # 2 columns, 1 row of images (each initially 300×300)
canvases = gui["canvas"]
imshow(canvases[1,1], testimage("lighthouse"))
imshow(canvases[1,2], testimage("mandrill"))
Gtk.showall(gui["window"])
Then I guess it might due to the version of some packages…
同时安装多个 package¶
用空格隔开,如
add Distributions Combinatorics MATLAB PyPlot LightGraphsFlows LightGraphs Clp Plots
内地镜像源¶
安装 PkgMirrors,则可以用中科大或者浙大的镜像源了
初次设置后,以后直接 using PkgMirrors 便切换到设置好的镜像,以后直接通过镜像源安装。
select text in the output pdf¶
Several weeks ago, the text, such as the label, title or legend, in the output pdf via savefig("xxx.pdf") can be selected, but recently I found that I cannot select the text in the pdf. The reason would be the version of the packages.
I had tried GRUtils, and GRUtils.savefig can output pdf to select the texts, so I think the reason is some changes in Plots package.
And I found that with Plots@0.27.0 in Julia1.0, I can select the text, but currently the newer Plots@1.2.0 cannot produce such pdf. To determine the change, maybe I need more effort to try different versions. But now I can downgrade the version to satisfy my requirement.
colorviews for images¶
check the help document by ?colorviews, and there are some examples to illustrate its usage, but here I add some examples that I used in my projects.
julia> colorview(RGB, rand(3,10,10))
10×10 reshape(reinterpret(RGB{Float64}, ::Array{Float64,3}), 10, 10) with eltype RGB{Float64}:
and we need to permutate the dims such that the first dim matches with RGB.
julia> colorview(RGB, permutedims(rand(10, 10, 3), (3, 1, 2)))
10×10 reshape(reinterpret(RGB{Float64}, ::Array{Float64,3}), 10, 10) with eltype RGB{Float64}:
we also can append the alpha channel to an image, such as
julia> colorview(RGBA, colorview(RGB, rand(3, 10, 10)), rand(10, 10))
10×10 mappedarray(RGBA{Float64}, ImageCore.extractchannels, reshape(reinterpret(RGB{Float64}, ::Array{Float64,3}), 10, 10), ::Array{Float64,2}) with eltype RGBA{Float64}:
but directly append the array does not work,
julia> colorview(RGBA, rand(3, 10, 10), rand(10, 10))
ERROR: DimensionMismatch("arrays do not all have the same axes (got (Base.OneTo(3), Base.OneTo(10), Base.OneTo(10)) and (Base.OneTo(10), Base.OneTo(10)))")
which should be replaced with
julia> colorview(RGBA, rand(4, 10, 10))
10×10 reshape(reinterpret(RGBA{Float64}, ::Array{Float64,3}), 10, 10) with eltype RGBA{Float64}:
To use RGB{N0f8}, the input array should be UInt8 not Int,
julia> colorview(RGB{N0f8}, Array{UInt8}(fill(1,3,10,10)))
10×10 reshape(reinterpret(RGB{N0f8}, view(::Array{UInt8,3}, [1, 2, 3], :, :)), 10, 10) with eltype RGB{Normed{UInt8,8}}:
return values of function¶
julia> versioninfo()
Julia Version 1.4.0
Commit b8e9a9ecc6 (2020-03-21 16:36 UTC)
Platform Info:
OS: Linux (x86_64-pc-linux-gnu)
CPU: Intel(R) Core(TM) i5-6300HQ CPU @ 2.30GHz
WORD_SIZE: 64
LIBM: libopenlibm
LLVM: libLLVM-8.0.1 (ORCJIT, skylake)
julia> function f(a, b, c, d)
return a, b, c, d
end
julia> a = f(1,2,3,4);
julia> a
(1, 2, 3, 4)
julia> a, b = f(1,2,3,4);
julia> a
1
julia> b
2
julia> a, b, c = f(1,2,3,4);
julia> a
1
julia> b
2
julia> c
3
String¶
padding zero on the left¶
For example, convert “1” to “001”,
julia> lpad(1, 3, '0')
"001"
>>> a = 1
>>> f"{a:03}"
'001'
check if substring¶
julia> occursin("tech", "techNotes")
true
>>> "tech" in "techNotes"
True
single/double quoting¶
julia> a = 'www'
ERROR: syntax: invalid character literal
julia> a = 'w'
'w': ASCII/Unicode U+0077 (category Ll: Letter, lowercase)
julia> a = "ww"
"ww"
refer to #105.
run command line¶
For example, to combine different figures,
run(`convert p1.png p2.png +append p12.png`)
but if there are two many figures to combine, the correct way is to keep the multiple figure name as an array,
figs = "p" .* string.(1:10) .* ".png"
run(`convert $figs +append pall.png`)
instead of trying to convert the array to a string,
# WRONG!!!
julia> figs = prod("p" .* string.(1:10) .* ".png ")
julia> run(`convert $figs +append pall.png`)
convert: unable to open image 'p1.png p2.png ... p10.png '
or writting all command as a string
# WRONG!!!
command = "convert " * prod("p" .* string.(1:10) .* ".png ") * "+append pall.png"
run(`$command`)
Special characters #{}()[]<>|&*?~; need to be quoted,
julia> run(`ls *.toml`)
ERROR: LoadError: parsing command `ls *.toml`: special characters "#{}()[]<>|&*?~;" must be quoted in commands
the quote is not just for the special character, which will fix the filename in the above example,
julia> run(`ls "*.toml"`)
ls: cannot access '*.toml': No such file or directory
but quoting is not enough,
julia> run(`"ls *.toml"`)
ERROR: IOError: could not spawn `'ls *.toml'`: no such file or directory (ENOENT)
the right way is
julia> run(`bash -c "ls *.toml"`)
Manifest.toml Project.toml
Process(`bash -c 'ls *.toml'`, ProcessExited(0))
refer to Quoting in shell commands - General Usage - JuliaLang
foreach¶
julia> foreach(readdir(".")) do filename
println(filename)
end
# or a more concise way
julia> foreach(println, readdir("."))
Regex¶
julia> split("juliaxxxjulia", "julia")
3-element Array{SubString{String},1}:
""
"xxx"
""
# ^ requires `Regex`
julia> split("juliaaaajulia", "^julia")
1-element Array{SubString{String},1}:
"juliaaaajulia"
julia> split("juliaxxxjulia", Regex("^julia"))
2-element Array{SubString{String},1}:
""
"xxxjulia"
julia> x = "xxx"
"xxx"
julia> split("juliaxxxjulia", Regex("^julia$x"))
2-element Array{SubString{String},1}:
""
"julia"
adopted from pattern = Regex("^"*pattern)
Underscores¶
julia> function ff()
return 1, 2, 3
end
ff (generic function with 1 method)
julia> _, _, a = ff()
(1, 2, 3)
julia> a
3
julia> _
ERROR: all-underscore identifier used as rvalue
where the formal definition for rvalue refers to Value categories